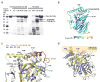- Journal List
- NIHPA Author Manuscripts
- PMC1431608
Structural mechanism for sterol sensing and transport by OSBP-related proteins
Abstract
Back to the Tutorial Index 1ZHW...Back to the Tutorial Index 1ZHT...The oxysterol binding protein (OSBP)-related proteins (ORPs) are conserved from yeast to man The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
OSBP was first discovered The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
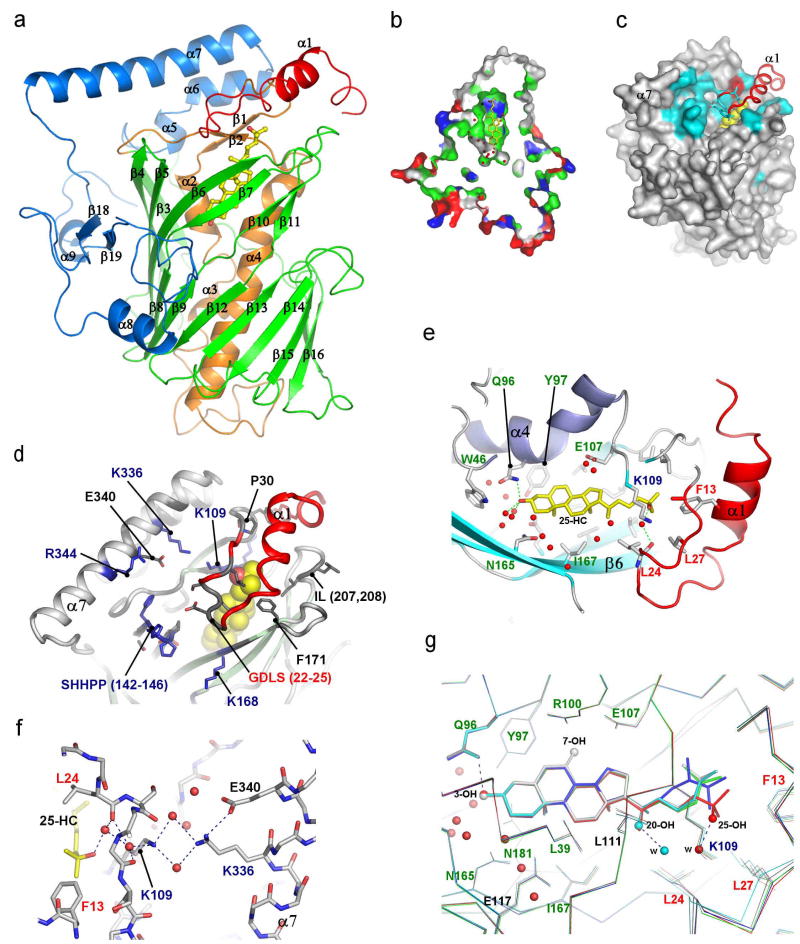
The structure of full-length Osh4 was determined by multiple isomorphous replacement and refined to a free R-factor of 23 % at 1.5 Å resolution (Supplementary Fig. 1; Supplementary Table 1). Osh4 is built around a central antiparallel β-sheet of 19 strands (residues 115–293) which form a nearly complete β-barrel (Fig. 1a, Supplementary Fig. 2). The strands of the barrel are pitched at ~45 degrees to its axis. The barrel has structural similarity to the large β-barrels of various bacterial outer membrane transporters The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
Sterols bind to Osh4 in the central tunnel of the β-barrel in a head down orientation. The 3-hydroxyl is buried at the bottom of the tunnel and the side-chain touches the inner surface of the lid (Fig. 1e). Although Osh4 has a novel fold, the burial of its ligands in a central hydrophobic tunnel is reminiscent of the structures of other lipid binding and transport proteins. The closest analogy is to the cholesterol-binding steroidogenic acute regulatory protein (StAR) transport (START) domain proteins MLN64 The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
The most striking aspects of oxysterol recognition by Osh4 are the absence of direct hydrogen bonds between sterol hydroxyl groups and side-chains of conserved amino acids, and the prominence of water-mediated interactions. The 3-hydroxyl of cholesterol, ergosterol, and oxysterols binds to two water molecules and to the side-chain of Gln 96. This Gln is part of a hydrated cluster of polar side-chains at the bottom of the tunnel (Fig. 1e, g). Trp 46, Tyr 97, Asn 165, and Gln 181 comprise the remainder of the polar cluster at the tunnel bottom, and form water-mediated interactions with the 3-hydroxyl. The 20- and 25-hydroxyl groups of the respective oxysterols interact with more or less ordered water molecules but not directly with the protein. The 7-hydroxyl of 7-hydroxycholesterol (7-HC) has no apparent hydrogen bonds to either protein or waters. The 25-hydroxyl is linked via two water molecules to the conserved Lys-109 side-chain, but other oxysterol hydroxyls are linked to less well-ordered waters. The lack of direct protein interaction with oxysterol hydroxyls in Osh4 contrasts sharply to the specific recognition of 24(S), 25 epoxycholesterol by a His-Trp pair in LXR The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
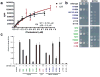
The absence of direct interactions between hydroxyl groups and the protein is difficult to reconcile with the concept that Osh4 is a specific effector of oxysterol signaling. We examined the relative affinities of Osh4 for cholesterol and oxysterols. Cholesterol binds to Osh4 with a Kd of 300 nM (Fig. 2a), as compared to a Kd of 55 nM for the tightest-binding oxysterol, 25-HC (Supplementary Fig. 5). Given the modest difference in affinities, and given that cholesterol is far more abundant than oxysterols in mammalian cells, we reasoned that cholesterol might be a physiological ligand for ORPs. We report elsewhere tests of this hypothesis that show Osh4 transports ergosterol, the yeast counterpart of cholesterol (Raychaudhuri, Im, Hurley & Prinz, in preparation).
In the bound conformation of Osh4, sterol ligands are inaccessible from the outside. Therefore, a conformational change is required for the uptake and release of ligands. The lid has some of the highest B-values (Fig. 3b) in the structure, suggestive of flexibility. Following limited proteolysis by trypsin in the absence of ligand, Osh4 is rapidly converted to a stable fragment beginning at residue 28 (Fig. 3a), which corresponds to the end of the lid region (Fig. 3b). Binding of 25-HC stabilizes this region against proteolysis (Fig. 3a). Sterol ligands stabilize the closed conformation of the lid via direct van der Waals interactions with Trp 10, Phe 13, and the highly conserved residues Leu 24, and Leu 27 (Fig. 1f, g). We were unable to crystallize full-length Osh4 in the absence of ligand, which we attribute to the flexibility of the lid region in the unliganded state. We engineered a lidless Osh4 in which residues 1–29 were deleted and the flexible surface loop 236–240 was replaced by an ectopic dipeptide sequence. Mutation of loop 236–240 does not affect cholesterol binding (data not shown). We determined the 2.5 Å resolution structure of this form of Osh4 in its apo conformation (Fig. 3c, d).
The apo structure undergoes a dramatic conformational change (2.9 Å r.m.s. for all common Cα positions) with respect to the complexes. The change is triggered when the ligand-lid interactions are lost. The movement of the lid away from the rest of the protein unlocks α7 and basic-residue containing loops from their complexed conformations. In the apo structure the tunnel is open and there is no longer any barrier to ligand exchange. Helix α7 pivots about its N-terminus such that its C-terminus moves 15 Å. The β1 strand and several loop regions near the tunnel opening move by up to 7 Å. The Cα of the conserved basic residue Lys109 moves 6 Å in this conformational change. These shifts result in the reorganization of the entire conserved basic cluster at the tunnel entrance (Fig. 3c). The apo conformation presents putative phosphate binding sites and a flattened, unobstructed surface surrounding the tunnel opening, none of which are present in the bound conformation (Fig. 3d).
We tested whether the conserved basic residues at the tunnel entrance, as well as structural core, lid, and sterol-binding residues were important for biological function by assessing their ability to complement a yeast strain in which all seven ORPs have either been deleted or are present as a temperature-sensitive allele The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
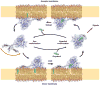
In the complexes, the ɛ-amino groups of Lys 109 and Lys 336 are ~3.6 Å away from each other in an unfavorable semi-buried environment (Fig. 1f). In the apo structure they move 8.7 Å apart. Two ordered sulfate ions are bound in the apo structure, one between the new positions of Lys 109 and Lys 336, and the other near the conserved His pair (Fig. 3c). We believe these sulfate ions mimic the roles of phospholipid phosphate groups because they bind in the position expected for the plane of the membrane surface during sterol exchange. The unfavorable geometry of the Lys 109-Lys 336 pair in the complex suggests that the closed structure is in a destabilized “tense” state. We propose that the open state is selectively promoted by the availability of phosphate groups at the membrane surface, which bind to these Lys only in the open state. Thus, these residues appear to function as an electrostatic spring-loaded lock that controls access to the tunnel. Based on these observations, we propose a sterol transfer and signaling cycle (Fig. 4).
OSBP is a cholesterol sensor that regulates phosphatase complexes The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
The lid and the basic cluster at the tunnel opening are conserved in all ORPs, while parts of the cholesterol binding site are not well conserved, suggesting that many ORPs might transport non-sterol ligands. The conservation of the lid and basic cluster lead us to believe that the transport cycle will be maintained throughout this protein family. Similarly, cholesterol-dependent signaling will require the same apparatus as transfer in order to take up cholesterol at membranes.
Methods
Protein expression and purification
DNA encoding Osh4 residues 2–434 was amplified by PCR and subcloned into the BamHI and XhoI sites of a pGEX-4T vector modified to contain a TEV protease recognition site. The fusion protein was expressed in E. coli BL21(DE3) overnight at 30 °C. The lysate was applied to glutathione sepharose. The GST was removed by digestion with TEV protease in the column and Osh4 was eluted with 500 mM NaCl and 50 mM NaH2PO4 (pH7.5). The protein was concentrated and then purified on a Superdex 200 column (Pharmacia) in 20 mM Tris-HCl (pH 8.0), 100 mM NaCl.
Crystallization
25-hydroxycholesterol was prepared in ethanol and added to 15 mg/ml protein at a final concentration of 1 mM. Crystals were grown by vapor-diffusion at 25 °C over a reservoir of 100 mM MES-NaOH (pH 6.5), 12% PEG 20000 over one week. Ergosterol, cholesterol, 7-hydroxy cholesterol, and 20-hydroxy cholesterol complex crystals were grown similarly. Crystals were cryoprotected in reservoir solution supplemented with 20% (v/v) glycerol and flash frozen under N2 gas at 95 K. Apo Osh4 crystals were grown in 100 mM MES-NaOH (pH 6.0) and 1.6 M ammonium sulfate using micro- and macro-seeding. The apo crystals were cryoprotected in reservoir solution containing 32% (v/v) glycerol. Structure determination is described in the supplementary methods.
Tryptic digestion and N-terminal amino acid sequencing
25-HC bound and apo-Osh4 were digested with increasing trypsin concentrations at 37°C for 2 hour. 25-HC bound Osh4 was prepared by adding 25-HC to the protein solution (0.3 mM) to a final concentration of 1 mM. 30 ug of each 25-HC bound Osh4 or apo-Osh4 was digested with increasing trypsin concentrations at 37°C for 2 hrs. From left to right samples were digested with 2.6, 0.45, 0.26, 0.052 μg/ml of trypsin. Fragments of the Osh4 protein prepared by the limited proteolysis using trypsin were separated by SDS-PAGE, transferred to PVDF membrane, and stained with Coomassie Blue R-250. Bands were cut out and the N-terminal amino acids were sequenced using a 492cLC Protein Sequencer (Applied Biosystems).
Ligand binding assay
For analysis of mutant Osh4 binding at a single concentration (Fig. 2a), [3H]-cholesterol and unlabeled cholesterol were mixed at a ratio of 1:9 to a final concentration of 10 μM in ethanol. Each reaction was carried out at room temperature in a final volume of 100 μl of 20 mM Tris pH 8.5, 30 mM NaCl, and 100 pmol Osh4. 15 pmol of cholesterol was added to the protein solution and incubated for 1 hr. Then 20 μl of Hightrap Q resin (70% (w/v)) (Amershan Biosciences) pre-equilibrated with Buffer A (20 mM Tris-HCl pH 8.5) was added and incubated for 5 min. The tubes were centrifuged at 13,000g for 1 min and washed with 1 ml of buffer A four times. The protein was eluted from the resin with 700 μl of buffer A containing 1 M NaCl. After centrifuging at 13,000g for 10 min, 200 μl of supernatant was taken and the radioactivity was measured in a liquid scintillation counter. In the low-stringency washing protocol (Fig. 2b), radiolabeled cholesterol (40 pmol) was dried under nitrogen and resuspended in 100 μL of 20 mM Tris, pH 8.5, 30 mM NaCl, 0.05% Triton-X-100. Purified Osh4p was added to a final concentration of 200nM and incubated at 30°C. After incubation for one hour, 15 μL of Hi-Trap Q resin (Amersham Pharmacia) was added and the samples were votexed and left at room temperature for five minutes. The resin was pelleted by spinning in a microcentrifuge at top speed for five minutes. After three washes with 20 mM Tris (pH 8.5), the protein was eluted from the resin with 500 μL of 20 mM Tris, pH 8.5, 1 M NaCl. After centrifugation, the radiolabeled cholesterol in the supernatant was determined by scintillation counting. To determine nonspecific binding, 500 μM unlabelled cholesterol was included in the incubation.
Strains and complementation analysis
Complementation analysis of was performed by introducing plasmids expressing mutant Osh4p into either CBY926 The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version. The following popper user interface control may not be accessible. Tab to the next button to revert the control to an accessible version.
Materials
Correspondence and requests for materials should be sent to JHH. Coordinates have been deposited with the Protein Data Bank with accession numbers 1ZHT, 1ZHB, 1ZHW, 1ZHX, 1ZHY, 1ZHZ, and 1ZI7.
Acknowledgments
We thank T. Levine, V. Bankaitis, and M. Brown for discussions and sharing unpublished data, N. DeAngelis for excellent technical assistance, R. Craigie for assistance with protein sequencing, J. Kim for advice on the early stages of this project, G. Miller and H. Shi for collecting synchrotron data, C. Beh and R. Scheckman for yeast strains, F. Dyda for maintaining the home x-ray facility, and the staff of beamline X25, NSLS and of SER-CAT, APS, ANL for assistance with data collection. Y. J. I. thanks S. H. Eom for mentoring and support. Y. J. I. was partly supported by the Korea Science and Engineering Foundation. Research carried out at the National Synchrotron Light Source, Brookhaven National Laboratory, is supported by the U.S. Department of Energy, Division of Materials Sciences and Division of Chemical Sciences under Contract No. DE-AC02-98CH10886. Use of the Advanced Photon Source was supported by the U. S. Department of Energy, Basic Energy Sciences, Office of Science, under Contract No.W-31-109-Eng-38.
References
Formats:
- Article |
- PubReader |
- ePub (beta) |
- PDF (3.8M)
- Sterol binding and membrane lipid attachment to the Osh4 protein of yeast.[J Phys Chem B. 2010]Rogaski BLim JB, Klauda JB, . J Phys Chem B. 2010 Oct 28; 114(42):13562-73.
- Lipid-regulated sterol transfer between closely apposed membranes by oxysterol-binding protein homologues.[J Cell Biol. 2009]Schulz TAChoi MG, Raychaudhuri S, Mears JA, Ghirlando R, Hinshaw JE, Prinz WA, . J Cell Biol. 2009 Dec 14; 187(6):889-903.
- The sterol-binding protein Kes1/Osh4p is a regulator of polarized exocytosis.[Traffic. 2011]Alfaro GJohansen J, Dighe SA, Duamel G, Kozminski KG, Beh CT, . Traffic. 2011 Nov; 12(11):1521-36. Epub 2011 Sep 12.
- Characteristics of oxysterol binding proteins.[Int Rev Cytol. 2008]Yan DOlkkonen VM, . Int Rev Cytol. 2008; 265:253-85.
- Emerging roles of the oxysterol-binding protein family in metabolism, transport, and signaling.[Cell Mol Life Sci. 2008]Fairn GDMcMaster CR, . Cell Mol Life Sci. 2008 Jan; 65(2):228-36.
- STRUCTURAL BASIS OF STEROL BINDING BY NPC2, A LYSOSOMAL PROTEIN DEFICIENT IN NIEMANN-PICK TYPE C2 DISEASE[The Journal of biological chem...]Xu S, Benoff B, Liou HL, Lobel P, Stock AM. The Journal of biological chemistry. 2007 Aug 10; 282(32)23525-23531
- The Yeast Oxysterol Binding Protein Kes1 Maintains Sphingolipid Levels[PLoS ONE. ]LeBlanc MA, Fairn GD, Russo SB, Czyz O, Zaremberg V, Cowart LA, McMaster CR. PLoS ONE. 8(4)e60485
- Cholesterol Modulates Cell Signaling and Protein Networking by Specifically Interacting with PDZ Domain-Containing Scaffold Proteins[Nature communications. 2012]Sheng R, Chen Y, Gee HY, Stec E, Melowic HR, Blatner NR, Tun MP, Kim Y, Källberg M, Fujiwara TK, Hong JH, Kim KP, Lu H, Kusumi A, Lee MG, Cho W. Nature communications. 2012 Dec 4; 31249
- Vesicle trafficking from a lipid perspective: Lipid regulation of exocytosis in Saccharomyces cerevisiae[Cellular Logistics. 2012]Johansen J, Ramanathan V, Beh CT. Cellular Logistics. 2012 Jul 1; 2(3)151-160
- Pathogen-Related Yeast (PRY) proteins and members of the CAP superfamily are secreted sterol-binding proteins[Proceedings of the National Ac...]Choudhary V, Schneiter R. Proceedings of the National Academy of Sciences of the United States of America. 2012 Oct 16; 109(42)16882-16887
- BioAssayBioAssayPubChem BioAssay links
- GeneGeneGene links
- GEO ProfilesGEO ProfilesRelated GEO records
- HomoloGeneHomoloGeneHomoloGene links
- OMIMOMIMOMIM record citing PubMed
- Pathways + GOPathways + GOPathways, annotations and biological systems (BioSystems) that cite the current article.
- ProteinProteinPublished protein sequences
- PubMedPubMedPubMed citations for these articles
- StructureStructurePublished 3D structures
- SubstanceSubstancePubChem Substance links
- TaxonomyTaxonomyRelated taxonomy entry
- Taxonomy TreeTaxonomy Tree
- Structural mechanism for sterol sensing and transport by OSBP-related proteinsStructural mechanism for sterol sensing and transport by OSBP-related proteinsNIHPA Author Manuscripts. 2005 September 1; 437(7055)154PMC
- Structural mechanism for sterol sensing and transport by OSBP-related proteins.Structural mechanism for sterol sensing and transport by OSBP-related proteins.Nature. 2005 Sep 1 ;437(7055):154-8.PubMed
- Structural basis for hydroxycholesterols as natural ligands of orphan nuclear re...Structural basis for hydroxycholesterols as natural ligands of orphan nuclear receptor RORgamma.Mol Endocrinol. 2010 May ;24(5):923-9. doi: 10.1210/me.2009-0507. Epub 2010 Mar 4 .PubMed
Your browsing activity is empty.
Activity recording is turned off.
See more...- cDNA cloning and expression of oxysterol-binding protein, an oligomer with a potential leucine zipper.[J Biol Chem. 1989]Dawson PA, Ridgway ND, Slaughter CA, Brown MS, Goldstein JLJ Biol Chem. 1989 Oct 5; 264(28):16798-803.
- Review Oxysterol binding proteins: in more than one place at one time?[Biochem Cell Biol. 2004]Olkkonen VM, Levine TPBiochem Cell Biol. 2004 Feb; 82(1):87-98.
- Overlapping functions of the yeast oxysterol-binding protein homologues.[Genetics. 2001]Beh CT, Cool L, Phillips J, Rine JGenetics. 2001 Mar; 157(3):1117-40.
- A role for yeast oxysterol-binding protein homologs in endocytosis and in the maintenance of intracellular sterol-lipid distribution.[J Cell Sci. 2004]Beh CT, Rine JJ Cell Sci. 2004 Jun 15; 117(Pt 14):2983-96.
- OSBP is a cholesterol-regulated scaffolding protein in control of ERK 1/2 activation.[Science. 2005]Wang PY, Weng J, Anderson RGScience. 2005 Mar 4; 307(5714):1472-6.
- Biological activity of some oxygenated sterols.[Science. 1978]Kandutsch AA, Chen HW, Heiniger HJScience. 1978 Aug 11; 201(4355):498-501.
- Correlation between oxysterol binding to a cytosolic binding protein and potency in the repression of hydroxymethylglutaryl coenzyme A reductase.[J Biol Chem. 1984]Taylor FR, Saucier SE, Shown EP, Parish EJ, Kandutsch AAJ Biol Chem. 1984 Oct 25; 259(20):12382-7.
- Review Oxysterols: modulators of cholesterol metabolism and other processes.[Physiol Rev. 2000]Schroepfer GJ JrPhysiol Rev. 2000 Jan; 80(1):361-554.
- cDNA cloning and expression of oxysterol-binding protein, an oligomer with a potential leucine zipper.[J Biol Chem. 1989]Dawson PA, Ridgway ND, Slaughter CA, Brown MS, Goldstein JLJ Biol Chem. 1989 Oct 5; 264(28):16798-803.
- Review Oxysterol binding proteins: in more than one place at one time?[Biochem Cell Biol. 2004]Olkkonen VM, Levine TPBiochem Cell Biol. 2004 Feb; 82(1):87-98.
- Overlapping functions of the yeast oxysterol-binding protein homologues.[Genetics. 2001]Beh CT, Cool L, Phillips J, Rine JGenetics. 2001 Mar; 157(3):1117-40.
- A role for yeast oxysterol-binding protein homologs in endocytosis and in the maintenance of intracellular sterol-lipid distribution.[J Cell Sci. 2004]Beh CT, Rine JJ Cell Sci. 2004 Jun 15; 117(Pt 14):2983-96.
- The pleckstrin homology domain of oxysterol-binding protein recognises a determinant specific to Golgi membranes.[Curr Biol. 1998]Levine TP, Munro SCurr Biol. 1998 Jun 18; 8(13):729-39.
- A conserved ER targeting motif in three families of lipid binding proteins and in Opi1p binds VAP.[EMBO J. 2003]Loewen CJ, Roy A, Levine TPEMBO J. 2003 May 1; 22(9):2025-35.
- Translocation of oxysterol binding protein to Golgi apparatus triggered by ligand binding.[J Cell Biol. 1992]Ridgway ND, Dawson PA, Ho YK, Brown MS, Goldstein JLJ Cell Biol. 1992 Jan; 116(2):307-19.
- Novel members of the human oxysterol-binding protein family bind phospholipids and regulate vesicle transport.[J Biol Chem. 2001]Xu Y, Liu Y, Ridgway ND, McMaster CRJ Biol Chem. 2001 May 25; 276(21):18407-14.
- Analysis of oxysterol binding protein homologue Kes1p function in regulation of Sec14p-dependent protein transport from the yeast Golgi complex.[J Cell Biol. 2002]Li X, Rivas MP, Fang M, Marchena J, Mehrotra B, Chaudhary A, Feng L, Prestwich GD, Bankaitis VAJ Cell Biol. 2002 Apr 1; 157(1):63-77.
- OSBP is a cholesterol-regulated scaffolding protein in control of ERK 1/2 activation.[Science. 2005]Wang PY, Weng J, Anderson RGScience. 2005 Mar 4; 307(5714):1472-6.
- Review Beta-barrel proteins from bacterial outer membranes: structure, function and refolding.[Curr Opin Struct Biol. 1999]Buchanan SKCurr Opin Struct Biol. 1999 Aug; 9(4):455-61.
- Dali: a network tool for protein structure comparison.[Trends Biochem Sci. 1995]Holm L, Sander CTrends Biochem Sci. 1995 Nov; 20(11):478-80.
- Analysis of oxysterol binding protein homologue Kes1p function in regulation of Sec14p-dependent protein transport from the yeast Golgi complex.[J Cell Biol. 2002]Li X, Rivas MP, Fang M, Marchena J, Mehrotra B, Chaudhary A, Feng L, Prestwich GD, Bankaitis VAJ Cell Biol. 2002 Apr 1; 157(1):63-77.
- Structure and lipid transport mechanism of a StAR-related domain.[Nat Struct Biol. 2000]Tsujishita Y, Hurley JHNat Struct Biol. 2000 May; 7(5):408-14.
- Crystal structure of the Mus musculus cholesterol-regulated START protein 4 (StarD4) containing a StAR-related lipid transfer domain.[Proc Natl Acad Sci U S A. 2002]Romanowski MJ, Soccio RE, Breslow JL, Burley SKProc Natl Acad Sci U S A. 2002 May 14; 99(10):6949-54.
- Structure of human phosphatidylcholine transfer protein in complex with its ligand.[Nat Struct Biol. 2002]Roderick SL, Chan WW, Agate DS, Olsen LR, Vetting MW, Rajashankar KR, Cohen DENat Struct Biol. 2002 Jul; 9(7):507-11.
- Structure-function analysis of human [corrected] phosphatidylinositol transfer protein alpha bound to phosphatidylinositol.[Structure. 2004]Tilley SJ, Skippen A, Murray-Rust J, Swigart PM, Stewart A, Morgan CP, Cockcroft S, McDonald NQStructure. 2004 Feb; 12(2):317-26.
- Structure of a multifunctional protein. Mammalian phosphatidylinositol transfer protein complexed with phosphatidylcholine.[J Biol Chem. 2001]Yoder MD, Thomas LM, Tremblay JM, Oliver RL, Yarbrough LR, Helmkamp GM JrJ Biol Chem. 2001 Mar 23; 276(12):9246-52.
- Crystal structure of the Saccharomyces cerevisiae phosphatidylinositol-transfer protein.[Nature. 1998]Sha B, Phillips SE, Bankaitis VA, Luo MNature. 1998 Jan 29; 391(6666):506-10.
- The molecular basis of vitamin E retention: structure of human alpha-tocopherol transfer protein.[J Mol Biol. 2003]Meier R, Tomizaki T, Schulze-Briese C, Baumann U, Stocker AJ Mol Biol. 2003 Aug 15; 331(3):725-34.
- Crystal structure of human alpha-tocopherol transfer protein bound to its ligand: implications for ataxia with vitamin E deficiency.[Proc Natl Acad Sci U S A. 2003]Min KC, Kovall RA, Hendrickson WAProc Natl Acad Sci U S A. 2003 Dec 9; 100(25):14713-8.
- Crystal structure of human GM2-activator protein with a novel beta-cup topology.[J Mol Biol. 2000]Wright CS, Li SC, Rastinejad FJ Mol Biol. 2000 Dec 1; 304(3):411-22.
- Structure of a cholesterol-binding protein deficient in Niemann-Pick type C2 disease.[Proc Natl Acad Sci U S A. 2003]Friedland N, Liou HL, Lobel P, Stock AMProc Natl Acad Sci U S A. 2003 Mar 4; 100(5):2512-7.
- Structure of sterol carrier protein 2 at 1.8 A resolution reveals a hydrophobic tunnel suitable for lipid binding.[Biochemistry. 2000]Choinowski T, Hauser H, Piontek KBiochemistry. 2000 Feb 29; 39(8):1897-902.
- NMR structure of the sterol carrier protein-2: implications for the biological role.[J Mol Biol. 2000]García FL, Szyperski T, Dyer JH, Choinowski T, Seedorf U, Hauser H, Wüthrich KJ Mol Biol. 2000 Jan 21; 295(3):595-603.
- Structural basis for glycosphingolipid transfer specificity.[Nature. 2004]Malinina L, Malakhova ML, Teplov A, Brown RE, Patel DJNature. 2004 Aug 26; 430(7003):1048-53.
- X-ray crystal structure of the liver X receptor beta ligand binding domain: regulation by a histidine-tryptophan switch.[J Biol Chem. 2003]Williams S, Bledsoe RK, Collins JL, Boggs S, Lambert MH, Miller AB, Moore J, McKee DD, Moore L, Nichols J, Parks D, Watson M, Wisely B, Willson TMJ Biol Chem. 2003 Jul 18; 278(29):27138-43.
- Overlapping functions of the yeast oxysterol-binding protein homologues.[Genetics. 2001]Beh CT, Cool L, Phillips J, Rine JGenetics. 2001 Mar; 157(3):1117-40.
- Analysis of oxysterol binding protein homologue Kes1p function in regulation of Sec14p-dependent protein transport from the yeast Golgi complex.[J Cell Biol. 2002]Li X, Rivas MP, Fang M, Marchena J, Mehrotra B, Chaudhary A, Feng L, Prestwich GD, Bankaitis VAJ Cell Biol. 2002 Apr 1; 157(1):63-77.
- OSBP is a cholesterol-regulated scaffolding protein in control of ERK 1/2 activation.[Science. 2005]Wang PY, Weng J, Anderson RGScience. 2005 Mar 4; 307(5714):1472-6.
- A role for yeast oxysterol-binding protein homologs in endocytosis and in the maintenance of intracellular sterol-lipid distribution.[J Cell Sci. 2004]Beh CT, Rine JJ Cell Sci. 2004 Jun 15; 117(Pt 14):2983-96.
- Analysis of oxysterol binding protein homologue Kes1p function in regulation of Sec14p-dependent protein transport from the yeast Golgi complex.[J Cell Biol. 2002]Li X, Rivas MP, Fang M, Marchena J, Mehrotra B, Chaudhary A, Feng L, Prestwich GD, Bankaitis VAJ Cell Biol. 2002 Apr 1; 157(1):63-77.
- OSBP is a cholesterol-regulated scaffolding protein in control of ERK 1/2 activation.[Science. 2005]Wang PY, Weng J, Anderson RGScience. 2005 Mar 4; 307(5714):1472-6.